A tutorial dedicated to the use of FROGS using its command line interface on the migale server is proposed here : https://tutorials.migale.inrae.fr/posts/frogs-16s/. From a public dataset (Illumina 16S V3-V4), you will be able to run the full FROGS pipeline to create a BIOM file containing the OTUs, their abundances and their taxonomic affiliations.
 Migale Bioinformatics Facility
Migale Bioinformatics Facility
We provide several services for scientists to deal with life sciences data
Our Services
Planned shutdown of Galaxy service
Access to the Galaxy instance will be unavailable from January 29th to February 2nd 2021.
Expected duration of interruption
Start: 29/01/2021 6pm
End (estimation) : 03/02/2021 8am
Users affected by the incident
- Users of the Galaxy instance
Cause
Update to version 20.09
Thank you for your understanding
"Bioinformatics by practicing" cycle 2021
 The 2021 training cycle is open
The 2021 training cycle is open
You can consult the program and the schedule here: https://migale.inra.fr/trainings
Tutorial
A new tutorial (only in french for now) about Migale usage :
- Access
- Storage
- Databanks
- Cluster usage , R
- Galaxy portal
We welcome all comments, by mail or through our annual satisfaction survey
Translation coming soon !
Rstudio on Migale
The Migale bioinformatics facility give acces to its users to a R environment (version 4.02) with more than 1000 packages installed.
For your development and prototyping activities with R, we now give you access to the RStudio development environment through a web interface at this adress :
https://rstudio.migale.inrae.fr
This RStudio is configured to use the R from Migale. It is linked to your work directories on the platform.
New publication: A Probiotic Mixture Induces Anxiolytic- and Antidepressive-Like Effects in Fischer and Maternally Deprived Long Evans Rats
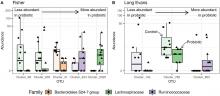
Mahendra and Olivier were sollicited by Valérie Daugé at MICALIS institute to analyze feces microbiota (V3-V4 regions of the 16S rDNA gene): α- and β-diversities and differential abundance analyses of OTUs.
Biological results:
Update R
R 4.0.2 is now the default version on migale
CHANGES IN R 4.0.2
UTILITIES:
* R CMD check skips vignette re-building (with a warning) if the VignetteBuilder package(s) are not available.
BUG FIXES:
* Paths with non-ASCII characters caused problems for package loading on Windows PR#17833.
* Using tcltk widgets no longer crashes R on Windows.
* source(*, echo=TRUE) no longer fails in some cases with empty lines; reported by Bill Dunlap in PR#17769.
* on.exit() now correctly matches named arguments, thanks to PR#1Planned shutdown of all services from September 29th to October 1st 2020
Access to the Migale bioinformatics facility will be unavailable from September 29th to October 1st 2020.
Expected duration of interruption
Start: 29/09/2020 10am
End (estimation) : 01/10/2020 2pm
Users affected by the incident
New article: Taxon Appearance From Extraction and Amplification Steps Demonstrates the Value of Multiple Controls in Tick Microbiota Analysis
A collaboration with Ecole Nationale Vétérinaire d’Alfort and INRAE teams led to the publication of this article. We showed that contaminant OTUs from sample laboratory processing steps can represent more than half the total sequence yield in sequencing runs, and lead to unreliable results when characterizing tick microbial communities. We thus strongly advise the routine use of negative controls in tick microbiota studies, and more generally in studies involving low biomass samples. You can access this article here.
New tools installation mode
From now on and as far as possible, we install our tools under the "conda" environment.
This allows us to avoid conflicts between the different dependencies of each tool.
We will gradually remove the "classic" "old" installations to keep only the "conda" versions. Beginning June 19th, all tools will be available only via Conda.
Help on how to use the tools under "conda" can be found here: https://migale.inra.fr/faq#migale-condalist